A cloud-based healthcare infrastructure for medical device integration: the bilirubinometer case study
Medical devices, generally, have been unique units, so that a typical hospital room in an intensive care ward hosts a big number of stand-alone devices, each one with its own user interface.
Annalisa Cenci, Daniele Liciotti,
Ilaria Ercoli and Primo Zingaretti
Department of Information Engineering
Universit`a Politecnica delle Marche
Ancona, Italy
{d.liciotti, i.ercoli, p.zingaretti}@univpm.it
Virgilio Paolo Carnielli
Neonatal Intensive Care Unit
Women’s and Children’s Hospital “G.Salesi”
Universit`a Politecnica delle Marche
Ancona, Italy
Abstract — Nowadays, these devices have some connection mechanisms, but the goal is to go beyond the simple connectivity that permits only the reading of the measured value on a computer monitor, to move towards a more complex device integration that offers the possibility to stream device data directly into patient electronic records, to create a database for further statistical investigations and to integrate the information derived from multiple medical devices into a single display. In this paper, we propose a method to connect and integrate into a cloud-based infrastructure the bilirubinometer, which is a typical Neonatal Intensive Care Unit biomedical device that measures the amount of bilirubin in infants’ blood, which is symptomatic of neonatal jaundice. After data extraction process from the instrument, the database is normalised according to a record format defined in XML language. Then, through Web Services of importation, data is transferred to the cloud database. It enables online sharing of encrypted healthcare data, reported in a standard language to allow a complete interoperability as well as a total security of shared data. Our proposed infrastructure allows the communication between different medical devices of a Neonatal Intensive Care Unit and permits the automation of the process of device data collection, transmission, storage, processing and availability for medical staff.
Keywords — Healthcare Infrastructure, Medical Device Integration, Cloud Computing, Database as a Service
I. INTRODUCTION
Medical devices generally are monolithic units so that a typical hospital room in an intensive care ward hosts a
number of stand-alone devices, each one with its own user interface, its own proprietary software, its own display and often its own built-in computer. Graphical user interfaces are all different and non-uniform and this can create confusion and misunderstandings among the healthcare personnel who has to use all the devices simultaneously.
2016 IEEE
![]()
Moreover, the devices are physically separated blocks and they can also be located on different bed sides so that the medical staff has to go from one device to another to use it, e.g., to see data on device display, to change device settings and to set up new vital parameters or new alarm values. In this way, the staff has to deal with different interfaces implemented on devices located in different places, which contributes to create mental confusion and poor organization and makes it more difficult for the physicians to rapidly take stock of the situation on the patient’s state of health. Furthermore, in a hospital ward there are many figures with different roles, e.g., the head physician, specialists in various fields and nurses, and each of them needs a tailored user interface suited to his own expertise and that shows him device data and information in the most suitable way to make him perform his job well and quickly.
Finally, around patient’s bed there are devices for the monitoring, the diagnosis and the treatment of different diseases and it would be appropriate to bring together data coming from all these devices to ensure that the physician could have a general idea about the patient’s health conditions in order to not neglect any important detail for the patient’s care. On the contrary, data are usually saved separately for each device and they are not always sent to the patient Electronic Health Record (EHR), so that it becomes difficult to search and find them, to create correlations between them and to make them available in real time to the clinician when he is visiting the patient, whereas this would bring great benefits to medical research [1].
Nowadays many marketed devices already have some connection mechanisms that use serial ports, Ethernet, IEEE 802.11 or Bluetooth wireless, but they are typically used to unidirectionally log data and events from these devices towards the computer, permitting the reading of the measured value on a PC display. However, this is no longer sufficient: as demand increases for better healthcare paradigms, it is evident that there is the need of having integrated and cooperating (interoperable) medical devices. Thus, the goal is to go beyond the simple connectivity to move towards a more complex device integration that offers the possibility to stream device data directly into patient electronic records, to create a database for further statistic investigations of data so collected and to integrate the information derived from multiple medical devices into a single customised display.
In this paper, we propose a cloud computing solution for the creation of a cloud infrastructure that exceeds the simple EHR. We created a cloud-based DBaaS (Databaseas-a-Service) with the abilities of storing and cooperatively
sharing medical data based on the HL7 message required for interoperability of medical information of different types. We create a database serving as a data consumer that allows the integration between device data, EHR data, personal patient’s data, patient’s medical history and so on. This permits to aggregate device data in one place, making easier the recorded data maintaining. This facilitates data searching in order to allow the doctor to make faster diagnosis, by having all relevant data available. Moreover, the identification of statistical trends and patterns is simplified by using data mining techniques that can lead to the discovery of correlations between collected data and illnesses, which otherwise would not have been possible to observe by clinicians [2] and that could be an useful tool to support a long-period management of a university hospital
[3].
The paper is organized as follows: next section describes the current scenario in the hospital ward taken as our case study (section II-A) and related works (section II-B); section III describes the proposed architecture; the evaluation of the system in practice is exposed in section IV and discuss and conclusions are presented in section V.
II. MOTIVATION
A. Current Scenario
The hospital ward that we take into consideration is the
Neonatal Intensive Care Unit (NICU) of the Women’s and Children’s Hospital ”G.Salesi” in Ancona, where preterm infants are taken into care. Preterm infants are babies born between the 22nd and 36th week of gestation [4] and they are infants with a high risk of mortality and morbidity. In fact, preterm birth affects the anatomical and functional development of all organs, inversely with gestational age and the degree of organs immaturity is a critical aspect for these babies, which leads them to have many health problems. For this reason, the primary physiological functions of these babies have to be continuously monitored and often infants have to undergo major treatments and therapies. Hence, infants’ cribs are surrounded by lots of devices for the monitoring, the diagnosis and the treatment of different diseases. As already explained, it is desirable that all these devices communicate between each other and with a unique database in order to create an integrated device network.
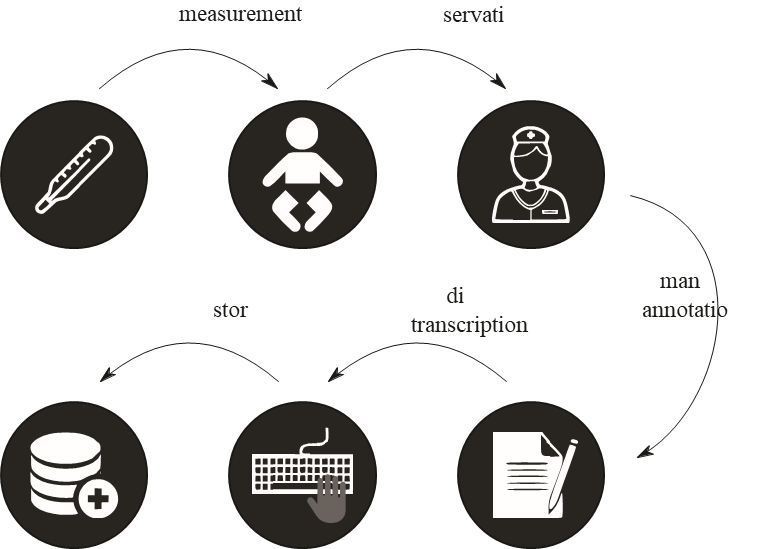
Nowadays, instead, the majority of these devices only provide the ability to read data on their digital displays. Thus, a member of the medical staff, generally the nurse in charge, has to collect infant’s data at bedside and to write them down to a paper medical chart on a regular basis. Then manual notes are regularly, but not immediately, transcripted by nurses in a PC of the Neonatal Unit. After that, data are transmitted to a local server that stores and organizes them together with the patient data collected in the EHR. Only at this moment, the medical staff can access data through an interface application. Focusing on the problem of infants’ vital data collection, distribution and processing, it is clear that current solution based on manual note writing is a slow and laborious process, susceptible to errors in the two phases of manual taking notes and of digital transcription on EHR. Moreover, there is an important delay between data generation and their availability and accessibility to medical staff. Fig. 1 shows this process based on manual taking notes.
We propose a method to make automated and faster this process that starts from crib data gathering and ends with the remote access to data by clinicians. We use medical communication protocols or proprietary software ability to automatically enable the communication and the data transfer between the devices and the unique database, which collects data from all medical devices.
We started this automation project by testing our proposed architecture with the transcutaneous bilirubinometer that is a typical biomedical device of a NICU. It makes a non-invasive measure of the amount of bilirubin in the blood of preterm infants, which is symptomatic of pathologic neonatal jaundice. The transcutaneous bilirubinometer is equipped with a sensor that, put in contact with the skin of the newborn, supplies as output the measurement of the transcutaneous bilirubin (TcB). Such non-invasive measurement is periodically repeated with a certain frequency (4-6-12 hours) that depends on the presence of evident jaundice and by the presence of risk factors. Measured TcB values are written down by the nurse on a spreadsheet and only later transcribed on the PC. Then, the doctor proceeds to the comparison of the measured TcB value with standard values established by neonatal jaundice guidelines [5]: the TcB value is visually compared with the correct curve selected depending on gestational age and on measurement time compared to birth hour. From this purely visual comparison and, therefore very subjective, the doctor decides if the value of TcB is in the physiological range or not. If it is, the nurse continues with the periodic TcB measurement; otherwise, if it is not, he proceeds with the measurement of the Total Serum Bilirubin (TSB) with a more invasive device that permits the analysis of a blood sample of the newborn heel. At this point, depending on the TSB value the nurse will return to perform the routine TcB measurements, or, in case of positive diagnosis, the physician will order to perform therapies such as phototherapy or exchange transfusion in more serious cases. As said before, since it is transcribed on PC only later, the TcB value has to be compared with the correspondent curve of the guidelines only visually through a paper chart. For this reason, this comparison is very subjective and prone to error.
Moreover, within the hospital ward there is more than one bilirubinometer and one of these is used in another room, where there are premature newborns in good health conditions that do not need special cares. In this case, measurements are not even transcribed from paper to EHR with the problem that if the newborn gets sick or gets worse and has to be moved to the NICU, they have to be recovered and transcribed on the PC in order to have the historical patient’s bilirubin values.
Thus, the automatic transmission of the measured values will eliminate these drawbacks and it will give the possibility of making comparisons with the standard curves of the guidelines in a direct, automatic and objective way and without the possibility of making mistakes.
The use of a single database, which also includes the personal data of the patient, e.g., the gestational age, the time and date of the birth, will allow the automatic selection of the guidelines curve with which the TcB measurement of a certain infant has to be compared and a simple algorithm will let the physician know if the TcB is outside the physiological range of values in an automatic way. Furthermore, in the future, thanks to the possibility of communication between different types of bilirubinometers (for serum and transcutaneous bilirubin measurements), when the measure of TcB will fall outside its physiological range, this will activate a signal that will be sent to the blood gas analyser and that will inform it in a direct way that it is necessary a TSB measurement. The possibility for medical devices to be data/control consumers could increase risks but can provide huge benefits including greater precision, reduction of human errors in repetitive tasks and cost reductions.
B. Related Works
The depicted scenario presents the opportunity to create an integrated cloud-based network that automates the process that begins with data collection from multiple devices and ends with information accessibility from medical staff. To achieve this goal we take advantage of diverse cut-edge technologies.
In the past, the healthcare information systems have been extensively implemented with traditional Management Information System (MIS) mode, but this does not comply with the current requirements of healthcare system. Lee et al. in [6] propose a service-oriented architecture (SOA)-based system that permits the integration of data from personal health devices and the creation of tailored users interfaces, but this work does not provide concurrency or constant and reliable service capabilities. Kulkarni and Ozturk in [7] implement a system called mPHASiS that leads to an end-to-end solution. Nonetheless, it is based on the Java programming language that makes the system less efficient for reasons of performance of Java Virtual Machine (JVM).
Our solution is based on the idea of cloud computing with which it is possible to provide customers with pervasive, affordable and on-demand services. Cloud computing owns the characteristics of parallel computing, distributed computing, and grid computing, but it improves the user experiences over the Internet compared to these. Lots of applications are moved towards a cloud platform because it is able to supply services for diverse application scenarios [8], [9]. Nowadays, also lots of works on medical information systems are based on cloud services [10], [11]. Cloud infrastructure can provide the flexibility and scalability of service-oriented systems unlike the transaction-oriented MIS mode.
Our proposed model provides support to ad hoc integration of diverse medical devices, distributed processing, and open accessibility. We propose a cloud-based infrastructure where medical devices are plugged in and begin to cooperate, i.e. to collect and transmit data. The computer resources achievable with this environment are set up to receive, store, process, and distribute the information. Our method permits the creation of an ad hoc integration network of different medical devices and of a distributed data processing.
The main requirement of our solution is that it has to quickly implement the methods to collect process and distribute infant’s vital data, from the crib to remote accessibility, by integrating all different medical devices [12], [13]. Moreover, it has to be flexible and extensible, i.e., it has to hold up diverse medical devices in different numbers that can be added to the system in different times [14]. Then, the proposed system has to guarantee the security and the safety of shared data, by ensuring strong access control and data encryption [15], [16]. To ensure data interoperability [17] it is necessary to realise data extraction process from various instruments, normalising the database according to a record format defined in the XML language and specifically to the logic related to language HL7 [18]. A database made with these characteristics enables online sharing of encrypted healthcare data, reported in a standard language, allowing a complete interoperability of shared data. The architecture has to guarantee also the system availability in any possible operational conditions, by ensuring its reliability [19] and it has to be scalable and compatible with the use in large healthcare environments and with the aggregation of different institutions [20]. In King et al. [21] they discuss about all these main problems and about their possible solutions with respect to the creation of a distributed architecture for the integration of medical devices in a hospital ward. They discuss about interoperability of data collected from devices, displays and databases of different vendors and about standardisation of device data streams and their reliability, security and safety, also in terms of privacy. However, the paper is not concentrated on concrete solutions, leaving room to other works for putting into practise the proposed ideas.
III. THE PROPOSED METHOD
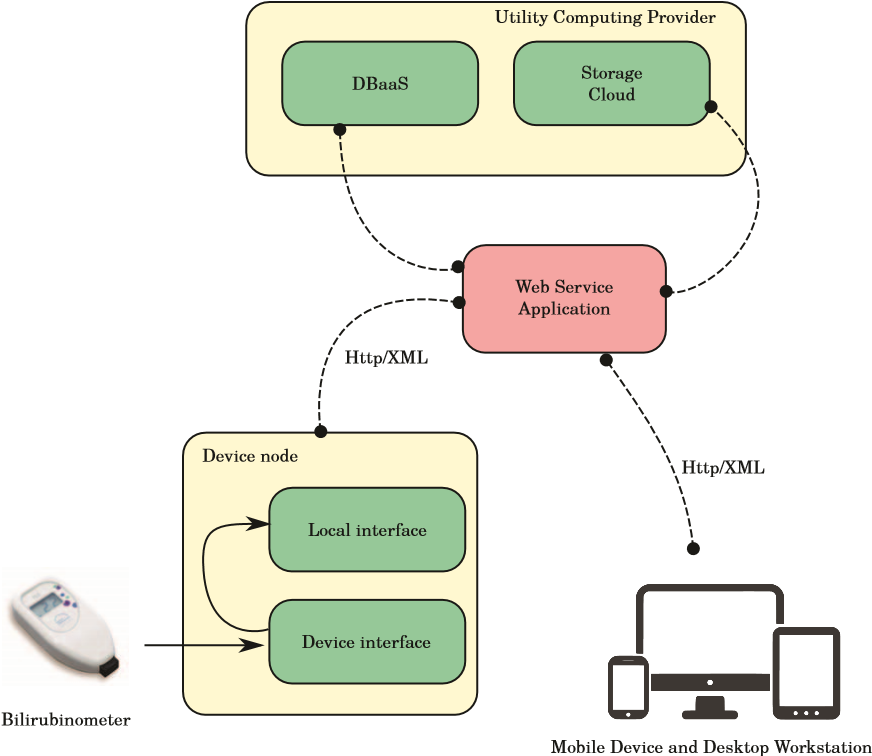
Fig. 2 shows our proposed system architecture that permits the developing and the deploying of the process automation of data collection, storage, processing and distribution through a medical device network that uses cloud computing.
At the preterm infant’s crib there are device nodes that are provided with hardware components such as serial port or USB port, forming the device interface, and with software to collect, encode, encrypt and transmit data through configurable communication channels to be stored, forming the local interface. Data are picked up through applications of extraction, specifically concerted for every different type of devices, which use medical communication protocols or proprietary software to enable the direct data communication and transfer. The process of extraction normalises data according to a record format defined in XML language and particularly according to the logic related to language HL7.
After extracting and standardising into XML, through Web Service of importation, data are carried to the cloud database, where they are aggregated.

The intrinsic interoperability, obtained by using different device types and vendors and language independent XML technologies and the widespread HTTP as a transport, implies that devices can communicate each other and with the EHR by using Web Services [22]. Web Services, in fact, are web application components that permit the interoperability between different technologies that work in a same network. These services hide their internal complexities such as data types and business logic from their users, but expose their programming interfaces and their locations using UDDI protocols. They make available a software interface, described in a format which can be automatically processed, which permits the interaction between other systems and the Web Service itself by activating the functions available on the interface (services or requests of remote procedures) through special messages of request. Such request messages are included in an ”envelope” that is formatted according to the standard XML, encapsulated and transported via Web protocol. In our case, the XML-based language for describing Web services application software interface is Web Services Description Language (WSDL), which is recommended by the World Wide Web Consortium (W3C). These interfaces, publicly exposed, guarantee the procedures of import data and access to the database by different authorized medical staff members for respective allowed functions. The XML-based protocol for accessing Web Services is Simple Object Access Protocol (SOAP), which is always recommended by W3C and the Web protocol on which SOAP operates is HTTP, the most commonly used and the only one that was standardised by the W3C.
Web services act like a broker between local medical devices and remote services. They are able to capture collected data from devices and to send them to suitable storage service hosted on cloud. They also allow the local storage for preprocessing, e.g., they allow data to be aggregated or simply analysed before the transfer to the main system.
On the other hand, Web Services work as an access point. Users through a web interface that acts like an application that works on Mobile Devices and Desktop Workstations, with which the hospital ward is equipped, can interact with data on the cloud database. Web Services receive requests from that interface to recover data from the cloud database.
The function of the Utility Computing Provider is to provide logical and physical infrastructure for data storage, processing and delivery services.
Its main components are the Platform front end interface, the Cloud Storage and the DBaaS (Database-as-a-Service). The platform front end interface is able to allow the management of the storage contents and to directly communicate with users. The interface can be a web client or a standalone application. The cloud storage is the model of data storage in which the digital data is stored in logical pools and it is based on highly virtualized infrastructure. Database-as-aservice [23] is one of cloud computing secondary service models and a key component of XaaS. This may be considered a subspecialty of the bigger Software-as-a-service model. In essence, DBaaS is a cloud database, which is a database that typically runs on a cloud computing platform. It is a cloud computing service model with which there is no need to physically launch a virtual machine instance for the database so that in such a configuration, application users do not have to install and maintain the database themselves. All of the administrative tasks, installation and maintenance are taken care of by the database service provider and application owners pay according to their usage.
Our project has been developed using as DBaaS the Amazon Relational Database (Amazon RDS) for MySQL that makes it easy to set up, operate, and scale MySQL deployments in the cloud. The use of this DBaaS provides to the system all the features that we have discussed in section II-B, which are the main requirements to obtain an efficient cloud-based healthcare infrastructure for medical device integration. It makes it easy to go from project conception to deployment. It is possible to access the capabilities of a production-ready relational database in minutes and without the need for infrastructure provisioning, and for installing and maintaining database software. It is possible to scale database compute and storage resources often with no downtime: the engine will automatically grow the size of database volume as database storage needs grow. It enhances reliability for critical production databases, including automated backups, database snapshots and automatic host replacement. For security reasons, it allows data encryption both during the transit, with the use of SSL, and during the cloud storage phase. Also backups, read replicas and snapshots are encrypted. Moreover, data are accessible only by authorized medical staff members by using strong authorization. There is also the possibility to run database instances in a virtual private network.
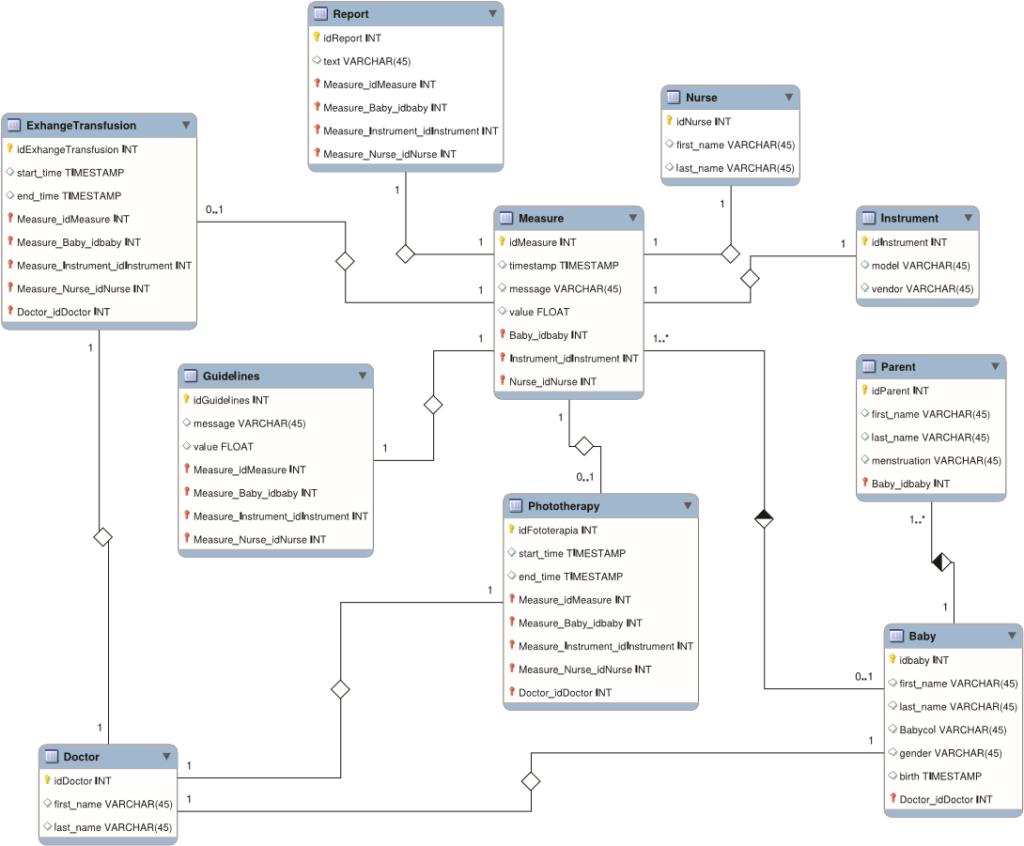
Fig. 3 shows the Enhanced Entity-Relationship (EER) diagram that illustrates the concepts we considered for modelling the portion of database with regard only to the case study of the transcutaneous bilirubinometer integration in the cloudbased healthcare infrastructure, but taking into care the future possibility of integrating into this device network also the blood gas analyser for the serum bilirubin measurement and, hence, of connecting the bilirubin measurement not only to guidelines, but also to phototherapy and exchange transfusion, as specified in section II-A.
Finally, we have created a Web Interface that works as an
application on Mobile Devices and Desktop Workstations. This service acts like a portal where medical staff members can access all available information. The user interface is built using the following architectures for different application areas: we use PHP as the server side scripting language; JavaScript as the client side scripting language and for the developing of the client side Web application it is used AngularJS, which is an infrastructure composed of a lot of functionalities. As already said, it is connected to the cloud database through WS-SOAP. It consists of a cross-platform solution that can be used with Windows, Linux and Mac, tablet Apple iOS, Android and Windows RT, while maintaining the same user interface. It is a simple and user-friendly interface that does not require special training. It is based on the paradigms of tablet and touch screen in order to minimize necessary interactions to access information and functions. Anyway, it is equipped with a user manual accessible through a click on the portal.
IV. EVALUATION OF THE SYSTEM AND RESULTS
TABLE I. DATABASE TRAFFIC
| Traffic | value per hour |
| Received | 471 M |
| Sent | 832 M |
TABLE II. DATABASE CONNECTIONS
| Connections | value per hour |
| Max contemporary connections | 578 |
| Failed attempts | 0 |
TABLE III. QUERY STATISTICS
| Queries | per hour |
| select | 386 k |
| insert | 36 |
| update | 27 |
| delete | 8 |
The HL7 message that we receive as bilirubinometer device output is composed of different segments: the message header segment (MSH), the patient identification segment (PID), the specimen model segment (SPC), the observation/result segment (OBX), the observation request segment (OBR) and the common order segment (ORC). We condense the most important segments fields in the following line: 000000000001|0000001|mg/dl|20160309165229|1.7| 20160309165250|3501198|20160309165250 − 0018|, where we find the most important output parameters that we use to create the database, as represented in EER diagram: the baby ID, the nurse ID, the unit of measurement, the measurement time, the measurement value, the message time, the device ID and the message ID.
We have conducted some initial studies by evaluating system performance in order to validate system usability. We have the first results from our system implementation regarding the integration of the transcutaneous bilirubinometer. They demonstrate that the solution provides good performance, by improving infants’ personal data availability by medical staff members and permitting data sharing with the bilirubinometer. Table I and table II present, respectively, the database performance about traffic and about connections, and table III shows the overall of the query statistics. The acquire results can be considered as indicative since the experiments are conducted in a real case scenario where the specific service is utilized in order to transmit and receive medical data.
As mentioned in the previous section, we also created a web platform where members of the hospital staff can log in via mobile or stationary devices (e.g., workstation, personal computer, tablet) through an authentication system (SSO Single Sign On) in order to increase the security of shared data. Each of them has a customised interface depending on their roles and on the operations that they can do. It is a highly configurable modular platform that can be easily adapted according to needs to be managed. In fact, we create the platform considering the possibility to expand it with new tools as new devices and new funcionalities are integrated in the healthcare infrastructure. The platform permits to browse personal data of the patient’s medical record and enter new patient data. Moreover, we implemented and tested the Bilirubin Tool, obviously connected with data coming from EHR, where physicians and nurses, once selected the involved infant, can view his historical values of bilirubin listed in a table or in a graphical way. Furthermore, once the TcB measurement is made, the value is displayed in real time on the PC monitor and it is automatically compared with the corresponding guidelines curve value, depending on the gestational age of the preterm infant and of the number of hours between birth and measurement time. After the automatic comparison, the system displays an alarm if measured bilirubin value falls outside of its physiological range. This completely eliminates the manual collecting laborious work and the resulting possibility of typing and comparison errors.
V. CONCLUSIONS AND FUTURE WORKS
In this paper, we presented an efficient cloud-based health infrastructure that permits the automation of the process of data collection, transmission, storage, processing and availability for NICU personnel. We have created a solid foundation for building a device network that allows data sharing between medical devices, as well as the communication with data of patient’s medical records in real-time and we tested it within the bilirubinometer case study. This allows the physician to have a general idea about the patient’s health conditions in order to not neglect any important detail for the patient’s care and to support him in his medical decisions. Moreover, the automatic transmission of the measured values from devices towards the cloud storage eliminates the drawbacks of manual collection work and the consequent possibility of typing errors. Data availability and accessibility, guaranteed from the web interface through Web services and ensured by strong authorization for only medical staff, facilitates the process of data delivery and visualisation. Furthermore, our infrastructure can provide the scalability necessary to support the adding of all medical devices that surrounded the neonatal crib for monitoring, diagnosis and treatment of different diseases, which is our next target. The idea is to obtain a single tailored display for each neonatal crib, where the clinician can access all patient’s data, e.g., he can visualise his medical records, all device data in real time, historical values, and where he can change device settings and set up new vital signs or new alarm values.
In the future, the possibility to share data and, especially, healthcare protocols, here investigated, will play a key role in the regional distribution of Neonatal Units. In fact, our method based on a flexible and extensible infrastructure is appropriate for the creation of a network that will include all the regional Neonatal Units. In this way, the system will facilitate the function of teleconsultation between specialists, enabling more effective collaboration between the peripheral structures and the central one, e.g., to better evaluate the conditions of a patient for which a transfer is requested or simply to unify healthcare protocols. The device integration in the infrastructure will make also possible to elaborate collected data in real time with specific algorithms of Machine Learning to assist physicians with clinical decision-making for improved healthcare in the view of Clinical Decision Support Systems.
ACKNOWLEDGMENT
The authors thank the healthcare personnel of the Neonatal Intensive Care Unit (Universita Politecnica delle Marche, An-` cona, Italy) for his precious collaboration and the continuous support to the work.
REFERENCES
[1] E. R. Millett, J. K. Quint, B. L. De Stavola, L. Smeeth, and S. L. Thomas, “Improved incidence estimates from linked versus stand-alone electronic health records,” Journal of Clinical Epidemiology, 2016.
[2] M. K. Obenshain, “Application of data mining techniques to healthcare data,” Infection Control & Hospital Epidemiology, vol. 25, no. 08, pp. 690–695, 2004.
[3] S. Tsumoto and S. Hirano, “Data mining in hospital information system for hospital management,” in 2009 ICME International Conference on Complex Medical Engineering, 2009.
[4] W. H. Organization et al., “Born too soon: the global action report on preterm birth,” 2012.
[5] C. Romagnoli, G. Barone, S. Pratesi, F. Raimondi, L. Capasso, E. Zecca, C. Dani et al., “Italian guidelines for management and treatment of hyperbilirubinaemia of newborn infants 35 weeks gestational age,” Italian journal of pediatrics, vol. 40, no. 1, p. 11, 2014.
[6] S.-H. Lee, J. H. Song, J.-H. Ye, H. J. Lee, B.-K. Yi, and I. K. Kim, “Soabased integrated pervasive personal health management system using phds,” in Pervasive Computing Technologies for Healthcare (PervasiveHealth), 2010 4th International Conference on-NO PERMISSIONS. IEEE, 2010, pp. 1–4.
[7] P. Kulkarni and Y. Ozturk, “mphasis: Mobile patient healthcare and sensor information system,” Journal of Network and Computer Applications, vol. 34, no. 1, pp. 402–417, 2011.
[8] J.-T. Hsu, S.-H. Hsieh, C.-C. Lo, C.-H. Hsu, P.-H. Cheng, S.-J. Chen, and F.-P. Lai, “Ubiquitous mobile personal health system based on cloud computing,” in TENCON 2011-2011 IEEE Region 10 Conference. IEEE, 2011, pp. 1387–1390.
[9] J. Biswas, J. Maniyeri, K. Gopalakrishnan, L. Shue, P. J. Eugene, H. N. Palit, F. Y. Siang, L. L. Seng, and L. Xiaorong, “Processing of wearable sensor data on the cloud-a step towards scaling of continuous monitoring of health and well-being,” in Engineering in Medicine and Biology Society (EMBC), 2010 Annual International Conference of the IEEE. IEEE, 2010, pp. 3860–3863.
[10] Z. Rashid, U. Farooq, J.-K. Jang, and S.-H. Park, “Cloud computing aware ubiquitous health care system,” in E-Health and Bioengineering Conference (EHB), 2011. IEEE, 2011, pp. 1–4.
[11] N. Botts, B. Thoms, A. Noamani, and T. A. Horan, “Cloud computing architectures for the underserved: Public health cyberinfrastructures through a network of healthatms,” in System Sciences (HICSS), 2010 43rd Hawaii International Conference on. IEEE, 2010, pp. 1–10.
[12] Q. Huang, L. Ye, M. Yu, F. Wu, and R. Liang, “Medical information integration based cloud computing,” in Network Computing and Information Security (NCIS), 2011 International Conference on, vol. 1. IEEE, 2011, pp. 79–83.
[13] Z. Chang, S. Mei, Z. Gu, J. Gu, L. Xia, S. Liang, and J. Lin, “Realization of integration and working procedure on digital hospital information system,” Computer Standards & Interfaces, vol. 25, no. 5, pp. 529– 537, 2003.
[14] C. O. Rolim, F. L. Koch, C. B. Westphall, J. Werner, A. Fracalossi, and G. S. Salvador, “A cloud computing solution for patient’s data collection in health care institutions,” in eHealth, Telemedicine, and Social Medicine, 2010. ETELEMED’10. Second International Conference on. IEEE, 2010, pp. 95–99.
[15] E. Frontoni, M. Baldi, P. Zingaretti, V. Landro, and P. Misericordia, “Security issues for data sharing and service interoperability in ehealth systems: the nu. sa. test bed,” in Security Technology (ICCST), 2014 International Carnahan Conference on. IEEE, 2014, pp. 1–6.
[16] R. Zhang and L. Liu, “Security models and requirements for healthcare application clouds,” in Cloud Computing (CLOUD), 2010 IEEE 3rd International Conference on. IEEE, 2010, pp. 268–275.
[17] L. Rossi, A. Belli, A. De Santis, C. Diamantini, E. Frontoni, E. Gambi, L. Palma, L. Pernini, P. Pierleoni, D. Potena et al., “Interoperability issues among smart home technological frameworks,” in Mechatronic and Embedded Systems and Applications (MESA), 2014 IEEE/ASME 10th International Conference on. IEEE, 2014, pp. 1–7.
[18] B. Orgun and J. Vu, “Hl7 ontology and mobile agents for interoperability in heterogeneous medical information systems,” Computers in biology and medicine, vol. 36, no. 7, pp. 817–836, 2006.
[19] M. Deng, M. Nalin, M. Petkovic, I. Baroni, and A. Marco, “To-´ wards trustworthy health platform cloud,” in Secure Data Management. Springer, 2012, pp. 162–175.
[20] C. He, X. Jin, Z. Zhao, and T. Xiang, “A cloud computing solution for hospital information system,” in Intelligent Computing and Intelligent Systems (ICIS), 2010 IEEE International Conference on, vol. 2. IEEE, 2010, pp. 517–520.
[21] A. L. King, S. Procter, D. Andresen, J. Hatcliff, S. Warren, W. Spees, R. P. Jetley, P. L. Jones, and S. Weininger, “An open test bed for medical device integration and coordination.” in ICSE Companion, 2009, pp. 141–151.
[22] J. Rao and X. Su, “A survey of automated web service composition methods,” in Semantic Web Services and Web Process Composition. Springer, 2004, pp. 43–54.
[23] H. Hacigum¨ us, B. Iyer, and S. Mehrotra, “Providing database as a¨ service,” in Data Engineering, 2002. Proceedings. 18th International Conference on. IEEE, 2002, pp. 29–38.